Models from Models
From statistics we know how to learn models from data: compute the likelihood \(L(model) = \mathrm{P}(data \mid model)\) and either maximize that or use it in a Bayesian update or something of the sort. This is a powerful mode of inference, but it's not the only type of inference possible, and I'd like to talk about a different type of inference today.
The wrong model
The normal statistical paradigm is what you might call model-based statistics: pick a model for your problem (a parameterized formula for \(P(data)\)); then use the observed data to learn the parameters. The way to screw this up is to pick the wrong model; for example, if your data are actually generated by a quadratic process, you will never fit a good line to the data.
How do you tell that you've got the wrong model? One way you can tell is if different subsets of data give different estimates. For linear data, you can make a good prediction even if you use just the left half (small \(x\)) of your data, and it won't be much different from the prediction with just the right half of the data:
>>> X, Y = genlin(1, 5) >>> linfit(X, Y) (0.99901650687168253, 4.9997591897673521) >>> linfit(X[X < .5], Y[X < .5]) (1.0367666060046121, 4.9938510330784167) >>> linfit(X[X > .5], Y[X > .5]) (1.050508547884262, 4.9575875290594702)
But if the data is quadratic, your best-fit line is very different if you restrict yourself to the left and right halves of your data set:
>>> X, Y = genquad(-1, 1, 1) >>> linfit(X, Y) (0.004178319672305573, 1.1615710546773379) >>> linfit(X[X > .5], Y[X > .5]) (-0.52382970878698176, 1.5607618274385056) >>> linfit(X[X < .5], Y[X < .5]) (0.50055285619314493, 1.0357115545536726)
I won't bother with confidence intervals here, but the agree: I should not have this big a difference for linear data unless I got very unlucky. A rough general heuristic emerges: if you can divide your data into two pieces that produce very different predictions, something's gone wrong.1 [1 This needs several qualifications. The first is that the pieces need to be equally weighted by your loss function; usually this means two pieces with roughly equally many points in both. The second is that you can't do this splitting after looking at the data; otherwise, you could pick “maximally confusing” points. In this post I'm going to talk about learning quadratics and lines, and I'll only ever use thresholds on \(x\) to split the data into pieces, so I should be safe.]
Upgrading models
Once we realize that our data is not linear, we must to switch to a different model; suppose we are lucky, and choose to switch to a quadratic model. How do we switch? One option is to re-learn the model from data. If the data is available, this is easy to do and gives us an optimal quadratic fit.
But what if the original data is now lost? Or, can we use the existing linear models to get a good quadratic fit more efficiently? For example, suppose values with \(x > ½\) were until recently impossible to observe. Then a good linear fit might exist for data with \(x < ½\), but the data itself may now be lost. Instead of learning models from data, can we learn models from models.
The broader philosophical2 [2 Scott Aaronson says that he’s interested in mathematics precisely because it is, the best way to tackle philosophical questions. I agree with all my heart.] question: when we realize our model was insufficiently rich, is any parameter-learning we did useless? Or can it be salvaged? It's often useful to have a model of part of the world—of a simple case, or of part of a phenomenon. But these models often conflict or contradict; how do humans handle using conflicting or contradictory models? How can we learn from our contradictory models? For me this ties into the idea of models as abstraction. A models abstract the data, but in my PL researcher hat I recognize that a good abstraction must itself be grounds for additional abstractions. We must be able to build models out of models.3 [3 I'd love to propose models-from-models as an exciting new field of statistics, but I don't have the research sense to know if that is wise. If it does become a field, I'm excited to read the developments.]
Upgrading for the linear equations
To upgrade from linear models to a quadratic model, it'll be helpful to define the problem more carefully. Let's suppose I have some data, and that the line \(y = a_{1i} x + a_{1i}\) is the best possible linear fit for the data points in \([l_i, r_i]\). I want to find the best quadratic fit \(c_2 x^2 + c_1 x + c_0\) that minimizes least-squares loss over my data.
Let me write \(f(x)\) for the quadratic I intend to learn, and \(g_i(x)\) for the $i$-th line.
\begin{align*} \mathcal{L} &= \sum_{(x, y)} (f(x) - y)^2 \\ &= \sum_{(x, y)} ((f(x) - g(x)) + (g(x) - y))^2 \\ &= \sum_{(x, y)} (f(x) - g(x))^2 + 2(f(x) - g(x))(g(x) - y) + (g(x) - y)^2 \end{align*}Note that by \(g\) here I mean whichever \(g\) applies to \(x\); let's assume that the intervals don't overlap.4 [4 Generalize all the way to knowing the distribution of \(x\) terms for each line if you want to weaken this condition.]
Now we can make a few simplifications. First, note that the last term does not depend on \(f\) so we can ignore it in our loss function.
\[ \mathcal{L} = \sum_{(x, y)} (f(x) - g(x))^2 + 2(f(x) - g(x))(g(x) - y) \]
Second, note that the first term does not care about our data—at least, it does not depend on \(y\). So let's suppose we know the distribution \(\mu(x)\) of \(x\) values. Then we can replace the sum over the data with an integral over the distribution of data:
\[ \mathcal{L} = \left(\int_\mathbb{R} (f(x) - g(x))^2 d\mu\right) + 2 \sum_{(x, y)} (f(x) - g(x)) (g(x) - y) \]
I want to minimize loss, so let's compute the derivative of the loss in \(c_k\). I differentiate under the integral sign.
\begin{align*} \partial_k\mathcal{L} &= \left(\int_{\mathbb{R}} 2(f(x) - g(x)) x^k d\mu\right) + 2 \sum_{(x, y)} x^k (g(x) - y) \\ &= 2 \left(\sum_i \sum_j (c_j - a_{ji}) \int_{l_i}^{r_i} x^{k + j} d\mu\right) + 2 \sum_{(x, y)} x^k (g(x) - y) \end{align*}We can call \(M^i\) the matrix whose entries are those integrals; then the first term reduces to:
\[ \partial\mathcal{L} = 2 \left(\sum_i M_i (c - a_i)\right) + 2 \sum_{(x, y)} x^k (g(x) - y) \]
I don't know how to make progress with the second term here. But one thought is that if we know that the data is actually on a quadratic, then the \(f\) we are learning is also an unbiased prediction for \(y\). So the derivative of \(\mathcal{L}\) is (expected to be)
\[ \partial\mathcal{L} = 2\left(\sum_i M_i (c - a_i)\right) + 2 \sum_{(x, y)} x^k (g(x) - f(x)) \]
Then the second term can be dealt with identically to the first, replacing the sum with an integral and then using these \(M_i\) matrices. Which gives the final equation
\[ \partial\mathcal{L} = 4 \sum_i M_i (c - a_i) \]
If we set this to zero (we are minimizing loss), we get
\begin{align*} 0 &= \sum_i M_i c - \sum_i M_i a_i \\ \left(\sum_i M_i\right) c &= \sum_i M_i a_i \\ c &= \left(\sum_i M_i \right)^{-1} \sum_i M_i a_i \end{align*}Note that all the matrices and vectors involved have size 3; this is incredibly computationally efficient, just linear in the number of lines. It's pretty much free as far as computation is concerned.5 [5 Not that least-squares fit is ever expensive…] Also note that if your lines partitioned the data, then \(\sum_i M_i\) is just the matrix of integrals over all of \(\mathbb{R}\), weighted by \(\mu\), so the inverse can be pre-computed and you don't need to do any sort of sum.
Testing this
I wrote a quick Python script to implement this inference idea. It's three lines of code in Numpy, and lets me play with this statistical tool; if you look at the script there are some quick tools to evaluate quality of fit, draw the resulting lines and quadratics, and so on. Here's an example with two lines and one hundred data points, linearly distributed between 0 and 1, with equation \(x^2 - x + 1\):
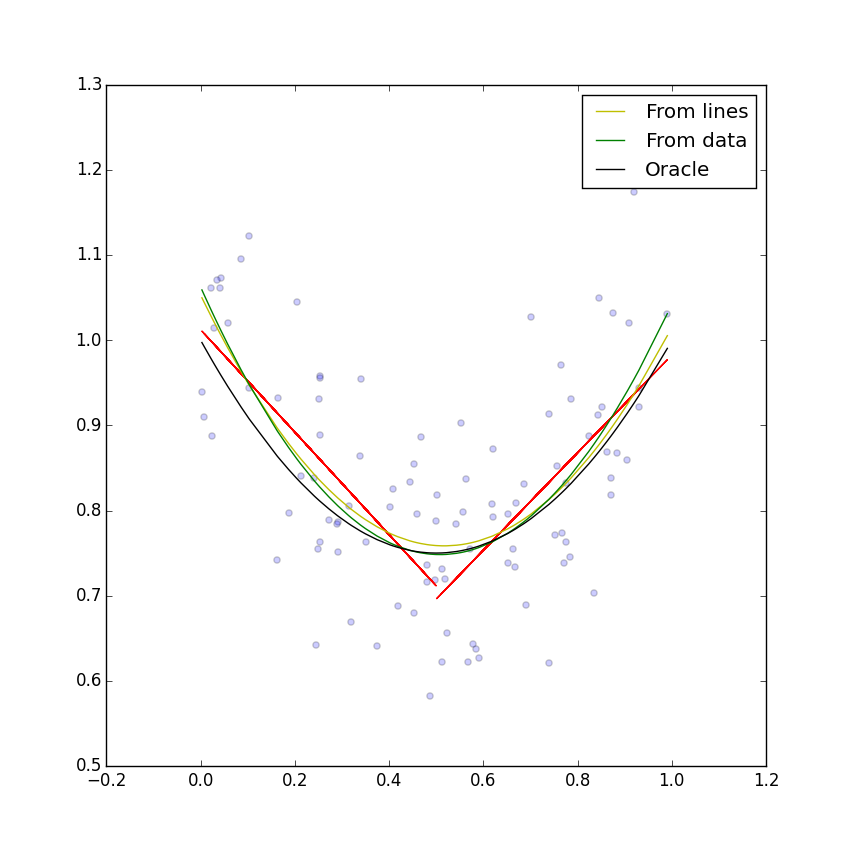
The fit is pretty good here—not obviously worse than the least-squares fit. I can compare both quadratic fits against the true equation, in terms of least-squares integral over \(x\) values (what you might call “true error”). The results are that the fit seems to be 9–17% worse than the least-squares fit (which is a provably optimal inference procedure).
It's surprising that just two lines work so well. Two “true” lines (best fit in terms of true error) are enough to derive the true quadratic, but more interestingly two “pretty close” lines are enough to derive “pretty close” quadratics. In a sense our abstraction of best linear fit is leaky: we do not go from a best linear fit to a best quadratic fit. But they don't leak much, either.
In fact, we can try out using a different number of lines, and see how that affects the quality of the fit. In this output, the first number is the number of lines used to get a quadratic fit, and the numbers in the tuple are how much worse the fit-from-lines was from a fit-from-data, in percentages (upper and lower 2σ confidence bounds over 1000 trials).
>>> [print(i, rtest([1, -1, 1], i)) for i in range(1, 10)] 1 (2424.2232066448178, 2745.1692124886849) 2 (8.7615854221890714, 17.518099247144178) 3 (0.35679928152396201, 5.3148503279066404) 4 (3.5842709136133521, 9.2025079794307203) 5 (3.7235846903315775, 9.766409268830941) 6 (5.8829552659600353, 13.541934172767256) 7 (13.183166348733337, 22.218958639745679) 8 (14.020041998179096, 23.377847111656919) 9 (17.779297477613071, 30.003014282232215)
What surprises me is that three lines looks like it provides the best estimator, just a few percentage points worse than a fit from data. This isn't true for all quadratics:
>>> [print(i, rtest([70, 1, 3], i)) for i in range(1, 10)] 1 (12300493.727961052, 13847923.571423735) 2 (60516.425614998727, 71116.021019622203) 3 (4713.2370783876122, 5643.5297291803408) 4 (1305.9569471418667, 1574.7237708896723) 5 (517.86376826549963, 634.22715636743749) 6 (280.95732263038087, 346.96845849937591) 7 (172.78419004566831, 216.65519744309796) 8 (125.36732912868595, 160.05879911302412) 9 (94.234909200141317, 123.50330186625089)
Here it looks like 9 lines isn't enough for this very non-linear parabola. I haven't really investigated the optimal number of lines much, but it seems like the next question.
The parabola coefficients are linear in the line coefficients. Since the coefficients of the best-fit line have error \(O(1/\sqrt{n})\) in the number \(n\) of input data points, the fit-from-lines coefficients will also have \(O(1/\sqrt{n})\) error asymptotically. As the number of data points increases, the percentage gap between the fit-from-lines and fit-from-data parabolas will converge to a constant. The linearity also means that, just like the fit-from-data quadratic, the fit-from-lines quadratic should be invariant under affine transformations.
So it sounds like fitting a parabola from lines is actually an acceptable way of fitting a parabola to data—a little worse than fitting it to data directly, but not too much worse either.
It's also fast. Finding a least squares fit for \(n\) data points of degree \(d\) takes time \(O(n d^2)\). The fit-from-lines requires \(O(n)\) time to compute the linear fits and, if you pre-compute \((\sum_i M_i)\) and its inverse, time \(O(d^2)\) to do the fit-to-lines. The improvement from \(O(n d^2)\) to \(O(n + d^2)\) is useless in practice, since it's extremely rare to do high-degree least-squares fits, but it's an interesting tid-bit in practice.
The quadratic from lines procedure I described can be generalized nicely into learning one general linear model by combining several general linear models, each defined on part of the data. But there's also a more general insight: to learn a model from models, you need to, instead of minimizing a sum over data points, minimize an sum over integrals, with each integral computing how well the learned model fits each sub-model. This could be a good avenue for extending any number of statistical tools to this model-from-models framework.
Footnotes:
This needs several qualifications. The first is that the pieces need to be equally weighted by your loss function; usually this means two pieces with roughly equally many points in both. The second is that you can't do this splitting after looking at the data; otherwise, you could pick “maximally confusing” points. In this post I'm going to talk about learning quadratics and lines, and I'll only ever use thresholds on \(x\) to split the data into pieces, so I should be safe.
Scott Aaronson says that he’s interested in mathematics precisely because it is, the best way to tackle philosophical questions. I agree with all my heart.
I'd love to propose models-from-models as an exciting new field of statistics, but I don't have the research sense to know if that is wise. If it does become a field, I'm excited to read the developments.
Generalize all the way to knowing the distribution of \(x\) terms for each line if you want to weaken this condition.
Not that least-squares fit is ever expensive…
